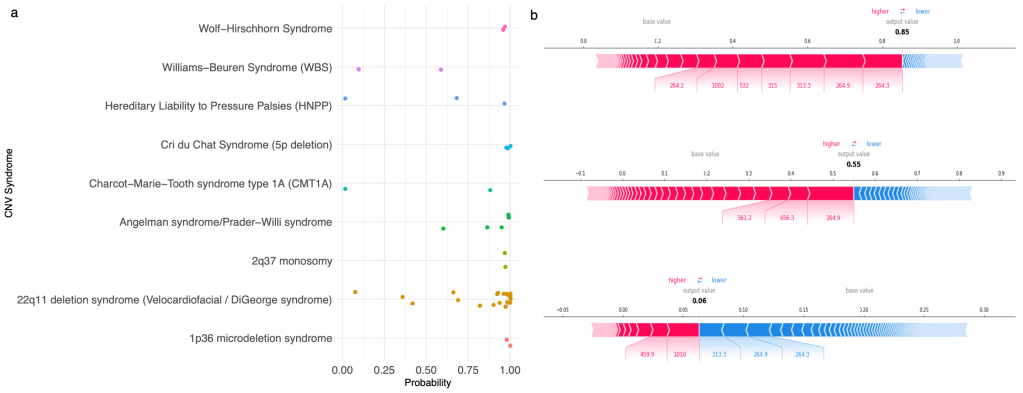
Identification of patients needing genetic testing or having a genetic disorder from EHR data
Genetic testing represents the primary means to diagnose an individual with a genetic disease. However, testing is provided in an inconsistent manner, potentially lengthening the diagnostic odysseys of patients who could benefit from them. We aim to utilize existing phenotype information in the form of diagnostic codes stored in the EHR to identify patients who could be helped by the administration of such a test. We hypothesize that individuals with genetic diseases tend to have a diverse constellation of rare phenotypes, and are therefore potentially identifiable without genetic information. To do this, we extract information from the set of individuals who have already received genetic tests, and train classifiers to identify patients with similar phenotypic profiles.
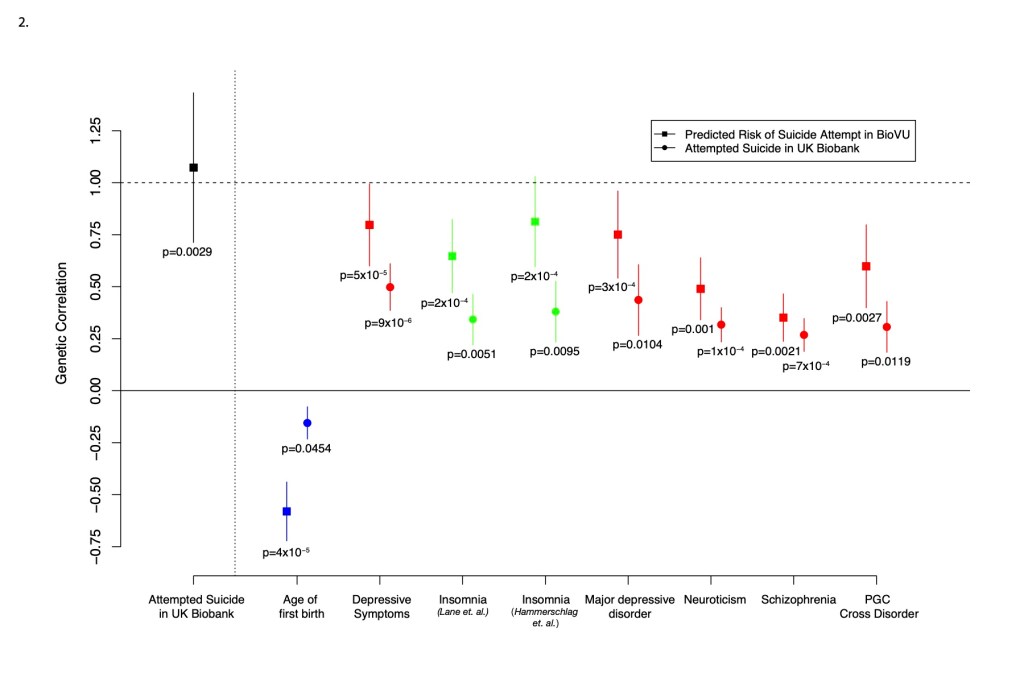
Genetic discovery using machine learning based phenotyping models in a large EHR
We are exploiting the exceptional resources in Vanderbilt’s biobank (bioVU) to capture novel phenotypes for studying behavioral health traits. For example, we are employing machine learning methods to predict individuals that are treatment resistant and using these predictions to perform pharmacogenomics at scale which has been a significant challenge in psychiatric research. This approach will enable us to integrate clinical and genetic data to probabilistically assign the best treatment for any given individual.
Capturing novel streams of behavioral health data from digital health applications and devices
Behavioral health information is rarely captured in clinical encounters and when it is often times it is not easily quantified. We are capturing novel streams of both objective and subjective behavioral health information from digital health applications and devices for incorporation in our clinical and genomic studies. We aim to study measures of behavioral health more broadly beyond disorders and utilize these measures to better characterize individuals across many dimensions for improved outcome prediction and treatment possibilities
Transcriptional consequences of structural variation in human brains
Structural variation (SV)– in particular a gain or loss of coding sequence – is known to contribute substantially to phenotypic diversity and disease. For example, in schizophrenia a burden of deletions and duplications contribute to disease risk. However, our understanding as to the biological underpinnings to this finding is minimal. In collaboration with the CommonMind Consortium we are mapping SV called from whole-genome sequencing to gene expression from RNA-sequencing to better understand the functional consequences of these events in 773 human post-mortem brains and relate these consequences to schizophrenia disease risk.

Intersecting genetics and pharmacology
Genes implicated by genomic studies represent possible therapeutic targets and can be used to inform drug design, drug repurposing and treatment efficacy. Recent large-scale GWAS of tens of thousands of cases and exome-sequencing of thousands of individuals have greatly increased the number genes and gene sets associated with risk to disease, however among the many important remaining questions is how these findings can inform and improve treatment. Our work focuses on intersecting the most recent findings of large-scale genetic studies with therapeutic information to identify potentially novel treatments at the population and individual level.
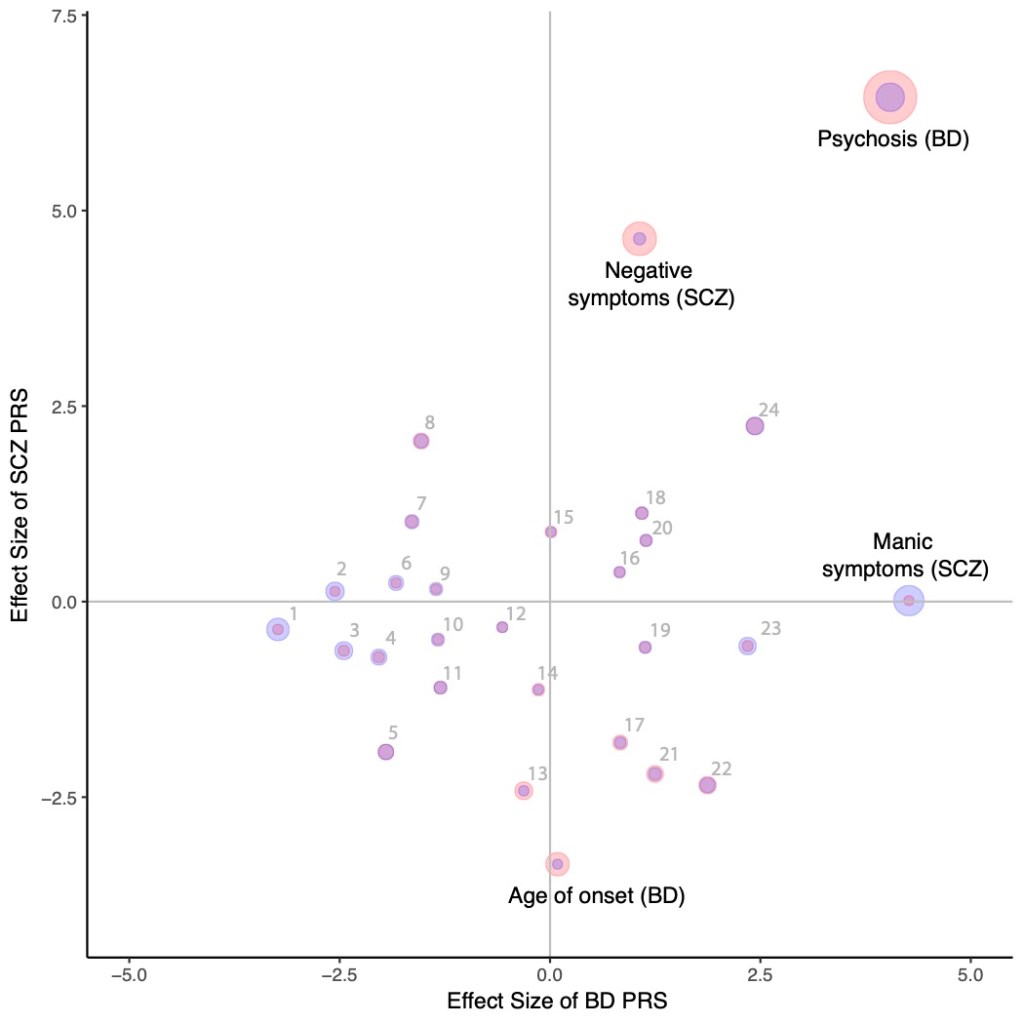
Large-scale genome-wide association studies
As part of the Psychiatric Genomics Consortium, we are seeking to gain insight into the fundamental genetic differences between related disorders like schizophrenia and bipolar disorder. We have previously identified a polygenic component that distinguishes these disorders but have yet to identify any particular genomic loci reaching genome-wide significance. We are currently compiling a much larger dataset to further this understanding. Additionally, we will incorporate our previous result showing a correlation between the bipolar genetic risk score and a clinical measure of mania in schizophrenia patients to build knowledge of molecular characterizations of disease. With larger samples and more phenotypic information we aim to improve risk prediction to be clinically useful in deciding appropriate treatment course for an individual with one of these disorders.